J.F. Bailey,1 B. Healy,1 H. Jianlin,2 L. Sherchand,3 S.L. Pradhan,4 T. Tsendsuren,5 J.M. Foggin,6 C. Gaillard,7 D. Steane,8 I. Zakharov 9 and D.G. Bradley1
1. Department of Genetics, Trinity College, Dublin 2, Ireland
2. Department of Animal Science, Gansu Agricultural University, Lanzhou 730070, Gansu, P.R. China
3. Livestock Production Division, Department of Livestock Services, Harihar
Bhawan, Pulchowk, Nepal
4. Resource Development Advisor, Nepal-Australia Community Resource Management Project, Kathmandu, Nepal
5. Institute of Biology, Academy of Sciences of Mongolia, Ulaan Baatar, Mongolia
6. Department of Biology, Arizona State University, Tempe, AZ 85287–1501
USA
7. Institute of Animal Breeding, University of Berne, Bremgarten-strasse 109a, CH-3012 Berne, Switzerland
8. FAO (Food and Agricultural Organization of the United Nations) Regional Office for Asia and the Pacific, 39 Phra Atit
Road, Bangkok 10200, Thailand
9. Vavilov Institute of General Genetics Russian Academy of Sciences, Gubkin str., 3, 117809 GSP-1, Moscow B-333, Russia
Yak (Bos grunniens) are members of the Artiodactyla, family Bovidae, genus Bos. Wild yak are first observed at Pleistocene levels of the fossil record. We believed that they, together with the closely related species of Bos taurus, B. indicus and Bison bison, resulted from a rapid radiation of the genus towards the end of the Miocene. Today domestic yak live a fragile existence in a harsh environment. Their fitness for this environment is vital to their survival and to the millions of pastoralists who depend upon them. Their wild counterparts are further marginalised to the most inhospitable areas of the Qinghai-Tibetan Plateau. Recent introduction of cattle (mainly B. taurus), in an attempt to improve stock, may threaten the integrity of the yak genetic make-up with the consequence of reducing fitness with respect to cold temperature and high altitude survival. Mitochondrial DNA (mtDNA) is ideally suited as a tool for studying population genetics because it has the unique features of maternal inheritance, a relatively fast rate of evolution and lack of recombination. Population history is traced back through the maternal line, which excludes the male genome and thus establishes a simple underlying pattern. It is possible to investigate the recent history of domestication due to the fast evolutionary rate of mtDNA. Lack of recombination between the mtDNA of yak and cattle is informative for the study of introgression. This paper discusses the genetic diversity of yak expressed in terms of sequence variations found in the displacement loop (D-loop) of mtDNA. A comparison has been made between these values and those found within other domestic species, primarily taurine cattle. An exact time for the domestication of yak is difficult to discern from archaeological data; here we estimate a time based upon a molecular clock of approximately 5000 years ago. Finally we shall feature the mtDNA introgression observed in yak populations.
Keywords: Cattle, introgression, mitochondrial DNA, sequence, variation, yak
Bos grunniens, B. mutus and Poephagus mutus are all species names, which have been assigned to yak. This variety of nomenclature reflects the uncertain relationship of the yak to other members of the Bovidae. The most widely used term is Bos grunniens and this will be used here. B. grunniens is grouped within a subfamily of the family Bovidae in the order Artiodactyla (even-toed ungulates) (Nowak 1991).
Establishing the phylogenetic position of B. grunniens within the Bovidae family has been problematic. Groves 1981 (cited in Janecek et al. 1996) suggested that the two species of bison were most closely related to yak, while alternative phylogenies place the Bison outside a clade containing all species of Bos. The recent application of molecular genetics to this field has clarified some, but not all, of these problems. In a cladistic analysis of mitochondrial ribosomal DNA from the family Bovidae, Gatesy et al. (1997) found that only that only in a few cases did the addition of morphological data overturn a clade favoured by the molecular data. Janecek et al. (1996) used the mitochondrial DNA (mtDNA) nucleotide sequence of the cytochrome oxidase II (Cox II) gene to infer phylogenetic relationships among the different taxa of the family Bovidae. They found a monophyletic clade of Bos/Bison and that the genus Bos is paraphyletic with respect to the genus Bison. Unfortunately the yak used in this study was a hybrid. Within the Bovini species it has been proposed that there is a correspondence between genetic distance and geographical origin. For example, yak are grouped closest with Bison in a principal co-ordinate analysis of amplified fragment length polymorphisms (AFLP) (Buntjer 1997, cited in Lenstra and Bradley 1999).
Yak inhabit areas of harsh conditions, generally living at altitudes of between 2000 and 5000 metre above sea level (masl). The average annual temperature in yak rearing areas is –8°C with yak being able to withstand temperatures of –40°C in the winter months (Shijan and Weisheng 1997). The average annual rainfall is 350–500 mm (Davaa 1997) and grasses in the area grow for only one-third of the year. Wild yak have no fixed territory and wander in search of food (Shijan and Weisheng 1997). Yak can cover an area of 200–300 km2 in search of food. Due to a shortage of grass during the winter, yak only consume an average of 40–60% of their daily fodder requirements. An individual animal can lose as much as 25–30% of its total body weight over the winter months (Davaa 1997).
We believed that B. grunniens were first domesticated on the Tibetan Plateau. According to Olsen (1990) the exact period when B. grunniens were domesticated is unknown, although archaeological evidence suggests this may have been some 5000 years ago (Cai and Wiener 1995; Ning et al. 1997). Yak are of great social and economic importance. Yak rearing is the major source of income for many nomadic and semi-nomadic people in China, Nepal, Bhutan, India, Pakistan, Mongolia, Russia, Tuva, Buryatia, Kirgizia and Tazakistan. Yak and yak products have a wide range of utility. Yak milk, a valuable source of protein for herders, is also used in the manufacture of cheese. The meat is popular because of its unique flavour. Yak hair and wool is used to make tents, ropes, bags and clothing. The livestock themselves are used to plough, carry produce and as a form of transport.
Phenotypic characters may help to distinguish different yak in different countries. For example, there are five breeds of yak found in India: Common, Bisonia, Bareback, White and Forehead Long Hair (Pal and Madan 1997). Cheng (1984) identified three types of yak in China: the Valley type, the Plateau Grassland type and the White yak, within these three types a number of breeds can be distinguished. As with European cattle certain breeds have been developed for particular traits. Common yak are most suitable for milk production while Bareback varieties should be used for meat. Brown yak tend to have finer undercoats and should be used for wool production (Pal and Madan 1997). Chinese yak, both white and black, are renowned for good quality meat with a distinctive flavour (Cheng 1984).
It has been proposed that yak breeding programmes will improve milk yields, meat quality, draft power, and wool productivity. Inbreeding remains a major obstacle for yak herders due to the difficulty of movement in these high altitude regions (Pal and Madan 1997). In an attempt to improve yak stock, programmes of hybridisation with cattle have taken place. Such programmes yield considerable benefits in production and may actually date back more than 3000 years (Shijan and Weisheng 1997). This approach has been treated with caution by those who are concerned that this may lead to a deterioration of the yak gene pool, which could result in a loss of fitness with respect to the cold temperature and high altitude survival (Tshering et al. 1997). Studies have demonstrated the advantage of mating wild and domestic yak (Shijan and Weisheng 1997). Such programmes pose little threat to the yak gene pool but may result in the production of some intractable animals.
A number of techniques have been used to study genetic diversity and molecular phylogeny. These include the assaying of protein polymorphism by protein immunology and protein electrophoresis, DNA hybridisation studies, restriction digests, repetitive DNA (e.g. microsatellites) and DNA sequencing of both nuclear and organelle DNA.
Mitochondria are organelles found within the cell. They are the sites for energy production during aerobic respiration. Each mammalian mitochondrion contains its own DNA, in the form of a covalently closed circular double stranded DNA molecule that contains 13 protein-coding genes, 2 ribosomal RNA (rRNA) genes and 22 transfer RNA (tRNA) genes (Gray 1989).
A number of characteristics make mtDNA a useful genetic marker; these include maternal inheritance, lack of recombination and fast rate of evolution.
No evidence for recombination has been found in interspecific hybrid cell lines (Clayton and Schadel 1997). Sperm mitochondria can enter the egg (Strauss 1999) but Avise (1991) thought that the absence of paternal mtDNA in the offspring is the result of an active mechanism in the oocyte, which removes paternal mtDNA.
MtDNA has a very rapid evolutionary rate. The rate of evolution of the mtDNA is different for different parts of the molecule. The tRNA and rRNA genes evolve approximately 100-fold quicker than their nuclear counterparts (Brown 1985; Avise 1991). The non-coding displacement loop (D-loop) evolves 5-times faster than the rest of the mtDNA. The silent site substitution rate in mammalian mtDNA has been estimated at 4.7 × 10–8 per site/year, i.e. 10-times higher than the rate for nuclear pseudogenes (Avise 1991).
A striking feature of mtDNA evolution is the ratio of transition substitutions to transversion substitutions, which is approximately 10:1 for the overall molecule (Gray 1989). This ratio is even more prominent in the D-loop where Bradley et al. (1996) observed a ratio of 57:1 for domestic cattle and the ratio for domestic yak was estimated at 36:1 (Bailey et al. in preparation). The ratio decreases as the divergence time increases due to multiple transitions at the same site. Explanations for the rapid rate of mitochondrial evolution include inefficiency of the repair mechanism (Cullinane and Bohr 1998) and relaxed functional constraint, partly due to the fact that mtDNA does not encode the proteins involved in its own replication, transcription and translation (Avise 1991). Other possible mechanisms are a high rate of turnover of mtDNA, a high exposure to oxidative damage and a relative lack of a recombinatorial mechanism (Avise 1991). It has also been suggested that mtDNA evolves more quickly than nuclear DNA due to a lack of associated histone proteins to the mtDNA (Gillespie 1986).
The D-loop region is non-coding and is involved in the control of replication and transcription (Clayton 1991). It is located between the tRNA genes for proline and phenylalanine. This region has been examined for nucleotide diversity and typed for some RFLP sites (Bailey et al. in preparation).
Bradley (2000) identified a classic 'double-headed broomstick' topology in an analysis of mtDNA sequence divergence for the four domestic ungulates: cattle, sheep, water buffalo and pig. In each case the mtDNA clusters into two distinct and divergent groups. The clusters are also geographically distributed and he argued that this pattern is indicative of a dual domestication pattern. Loftus et al. (1994) first noted a geographically distinct mtDNA clusters in cattle. One of their clades contained all the Indian cattle sequences, while the other contained African and European cattle sequences. They concluded that two separate domestication expansion events had occurred approximately 10 thousand years ago each from a highly divergent lineage, which separated approximately 0.2 to 1 MYA. The pattern of the domestic yak populations, taken from China, Bhutan, Nepal and Mongolia, is also a double-headed broomstick (Bailey et al. in preparation). This suggests that yak have undergone a domestication process similar to other ungulates. This dual domestication pattern differs from the pattern detected in other domesticates, such as horse and dog, which are known to have multiple domestication events.
Bradley (2000) noted that in most of the domestic populations a geographical distribution of haplogroups is observed. There are some exceptions to this observation: for example sheep. It has been suggested that secondary mixing has occurred in sheep populations because these animals are easily transported and may have been preferentially used in trade.
In yak mtDNA, two divergent haplogroups can be identified—Y1 and Y2. There does not appear to be a significant difference between the Chinese, Bhutanese, Nepalese and Mongolian yak (Bailey et al. in preparation). Yak have been referred to as the 'ships of the plateau' (Wenxiu 1997) and movement across the mountainous regions is only possible using yak. Such movements could act to reduce the amount of geographical partitioning observed, resulting in the pattern observed in Figure1.
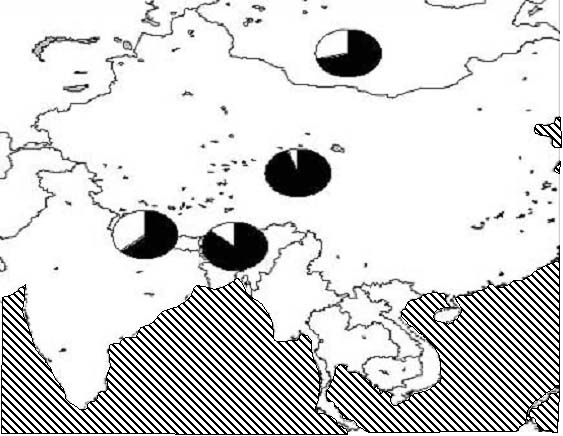
Data: Bailey et al. (in preparation).
Figure 1. A distribution map of the Y1 and Y2 haplogroups in samples from China, Bhutan, Nepal and Mongolia, where the shaded area of the pie is Y1 and the non-shaded area is Y2.
Molecular estimated dates for yak domestication, based on a molecular clock and subject to large errors, are roughly comparable to known archaeological data, which suggest a putative domestication date approximately 5000 years ago (Bailey et al. in preparation).
Nucleotide diversity values have been calculated for both the haplogroups Y1 and Y2 and for all Y (both 1 and 2). Diversity values can differ dramatically between domestic and wild populations. Therefore it is valuable to make a comparison between the nucleotide diversities found in yak and those found in another domestic species. Figure 2 is a comparison of the control region nucleotide diversities of the yak population and a box plot of the values found in 19 European cattle breeds. It clearly shows that the variability found in yak is of the same order as that found in European cattle. The similarity of diversity levels reflects the domestic history of these species. The total nucleotide variation, for both yak haplogroups, is highest because it reflects the whole mtDNA domestic pool from both centres of domestication. The Y2 is only based on a small number of individuals and is subject to change with future studies. The nucleotide diversity for Y1 is slightly lower than the median value for cattle. This relationship reflects the relatively recent domestication of yak approximately 5 thousand years ago as compared with the domestication of cattle (estimated to be as long as 10 thousand years ago).
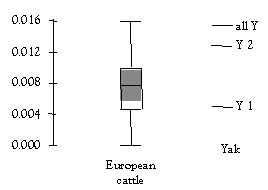
Data: Bailey et al. (in preparation). A boxplot of distribution of nucleotide diversities for 19 extant European breeds (MacHugh et al. 1999) is provided for comparison. Where the length of the box is interquartile range of the European nucleotide diversity and the horizontal line within it the median. The vertical lines outside the box extend to the smallest and largest observations and the shaded region the standard deviation.
Figure 2. Yak control region nucleotide diversities of all sequences studied and for the individual haplogroups Y1, Y2.
Bailey et al. (in preparation) note that some haplotypes are shared between animals from different populations. Investigations of mtDNA haplotypes by RFLP analysis (Jianlin 1997; Tu et al. 1998) also indicate that mtDNA haplotypes are shared across different Chinese yak breeds. Similarly, Tu et al. (1996) found that there were no significant genetic differences among the various yak populations in China at the level of protein polymorphism. Interestingly, haplotypes sharing across breeds and country groups is also observed within European cattle (Loftus et al. 1994; Bailey et al. 1996; Bradley et al. 1996).
MtDNA is not a useful tool for investigating domestic animals at the breed level because of the haplotype sharing between breeds for example as observed in both European cattle and yak populations. It would therefore be misguided to base yak breeding programmes on mtDNA surveys without additional knowledge of nuclear DNA. When microsatellite studies were made of 20 loci in European cattle it then became possible to observe resolution at the level of the breed (MacHugh et al. 1998). A similar study of both autosomal and sex linked microsatellites in yak should yield genetic structuring at the breed level. Such a study would be very informative for assisting with future breeding programmes.
Less than 1% cattle mtDNA was observed in a survey of yak mtDNA types (Bailey et al. in preparation). Whilst this is encouraging for those who are concerned about the effects of introgression (hybridisation with cattle) on yak it is important to note that mtDNA can remain unperturbed even when high levels of introgression are observed at the nuclear level (MacHugh 1997). It is therefore essential that a microsatellite study be conducted to allow assessment of introgression and facilitate yak-breeding programmes.
In summary the genetic investigation of yak mtDNA reveals a pattern consistent with other domestic ungulate species and suggests a dual domestication of these animals. The lack of phylogeographic distribution between yak populations is similar to the situation observed in sheep and probably reflects the mobile nature of yak. The genetic diversity observed in B. grunniens populations is comparable to levels found in European cattle populations. MtDNA is useful for examining the history of domestication within ungulate populations. It is essential to examine yak microsatellite markers to study the genetic integrity at the level of the breed.
Avise J.C. 1991. Ten unorthodox perspectives on evolution prompted by comparative population findings on mitochondrial DNA. Annual Review of Genetics 25:45–69.
Bailey J.F. (in preparation). Mitochondrial DNA variation in domestic yak populations.
Bailey J.F., Richards M.B., Macaulay V.A., Colson I.B., James I.T., Bradley D.G., Hedges R.E.M. and Sykes B.C. 1996. Ancient DNA suggests a recent expansion of European cattle from a diverse wild progenitor species. Proceedings of the Royal Society of London (series B) 263:1467–1473.
Bradley D.G. 2000. Mitochondrial DNA diversity and the origins of domestic livestock. In: Renfrew C. and Boyle K. (eds), Archaeogenetics: DNA and the population prehistory of Europe. MacDonald Institute for Archaeological Research, Cambridge, UK. pp. 315–320.
Bradley D.G., MacHugh D.E., Cunningham P. and Loftus R.T. 1996. Mitochondrial diversity and the origins of African and European cattle. Proceedings of the National Academy of Sciences (USA) 93:5131–5135.
Brown W. 1985. The mitochondrial genome of animals. In: MacIntyre R.J. (ed), Molecular Evolutionary Genetics. Plenum, New York, USA. pp. 95–130.
Buntjer J.B. 1997. DNA repeats in the vertebrate genome as probes in phylogeny and species identification. Academic thesis, Utrecht University. 130 pp.
Cai L. and Wiener G. 1995. The yak. FAO (Food and Agricultural Organization of the United Nations) Regional Office for Asia and the Pacific, Bangkok, Thailand. 237 pp.
Cheng P. 1984. Livestock breeds of China. Animal Production and Health Paper. FAO (Food and Agriculture Organization of the United Nations), Rome, Italy. pp. 96–108.
Clayton D.A. 1991. Replication and transcription of vertebrate mitochondrial DNA. Annual Review of Cell Biology 7:453–478.
Clayton D.A. and Schadel G.S. 1997. Mitochondrial DNA maintenance in vertebrates. Annual Review of Biochemistry 66:409–435.
Cullinane C. and Bohr V.A. 1998. DNA interstrand cross-links induced by psoralen are not repaired in mammalian Mitochondria. Cancer Research 58(7):1400–1404.
Davaa M. 1997. Conservation and management of yak genetic diversity in Mongolia. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 41–46.
Gatesy J., Amato G., Vrba E., Schaller G. and DeSalle R. 1997. A cladistic analysis of Mitochondrial ribosomal DNA from the Bovidae. Molecular Phylogenetics and Evolution 7(3):703–719.
Gillespie J.H. 1986. Variability of the evolutionary rates of DNA. Genetics 113:1077–1091.
Gray M.W. 1989. Origin and evolution of mitochondrial DNA. Annual Review of Cell Biology 5:25–50.
Groves C.P. 1981. Systematic relationships in the Bovini (Artiodactyla, Bovidae). Zeitschrift für Zoologie und Systematische Evolution 19:264–278.
Janecek L.L., Honeycutt R.L., Adkins R.M. and Davis S.K. 1996. Mitochondrial sequences and the molecular systematics of the Artiodactyl Subfamily Bovinae. Molecular Phylogenetics and Evolution 6(1):107–119.
Jianlin H. 1997. Yak genetic resources in China: evaluation of chromosome, protein and mtDNA polymorphism. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 175–183.
Lenstra J.A. and Bradley D.G. 1999. Systematics and phylogeny of cattle. In: Fries R. and Ruvinsky A. (eds), Genetics of Cattle. CAB (Commonwealth Agricultural Bureau) International, Wallingford, Oxon, UK. pp. 1–14.
Loftus R.T., MacHugh D.E., Bradley D.G., Sharp P.M. and Cunningham P. 1994. Evidence for two independent domestications of cattle. Proceedings of the National Academy of Sciences (USA) 91:2757–2761.
MacHugh D.E., Shriver M.D., Loftus R.T., Cunningham P. and Bradley D.G. 1997. Microsatellite DNA variation and the evolution, domestication and biogeography of taurine and zebu cattle (Bos taurus and Bos indicus). Genetics 146:1071–1085.
MacHugh D.E., Loftus R.T., Cunningham P. and Bradley D.G. 1998. Genetic structure of seven European cattle breeds assessed using 20 microsatellite markers. Animal Genetics 29:333–340.
Ning L., Jinge G. and Aireti. 1997. Yak in Xinjiang. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 115–122.
Nowak R.M. 1991. Walker's Mammals of the World. John Hopkins University Press, Baltimore. USA. 1407 pp.
Olsen S. 1990. Fossil ancestry of the yak, its agricultural significance and its domestication in Tibet. Proceedings of the National Academy of Sciences, Philadelphia, USA. 142:73–100.
Pal R.N. and Madan M.L. 1997. Yak production in India. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 29–39.
Shijan G. and Weisheng C. 1997. The situation of yak in China. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 25–28.
Strauss E. 1999. Can mitochondrial clocks keep time? Science 283:1435–1438.
Tshering L., Gyamtsho P. and Gyeltshen T. 1997. Yak in Bhutan. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 13–24.
Tu Z.C., Zhang Y.P. and Qiu H. 1996. Genetic diversity and divergence in Chinese yak (Bos grunniens) populations inferred from blood protein electrophoresis. Biochemical Genetics 35(1/2):13–16.
Tu Z.C., Zhang Y.P. and Qui H. 1998. Mitochondrial DNA polymorphism and genetic diversity in Chinese yak. Acta Genetica Sinica 25(3):205–212.
Wenxiu H. 1997. Conservation and management of yak genetic diversity in the Tibetan Autonomous Region. In: Miller D.G., Craig S.R. and Rana G.M. (eds), Proceedings of a workshop on conservation and management of yak genetic diversity held at ICIMOD, Kathmandu, Nepal, 29–31 October 1996. ICIMOD (International Centre for Integrated Mountain Development), Kathmandu, Nepal. pp. 93–104.