Q. Xuebin1,2 and H. Jianlin2
1. State Key Laboratory of Arid Agroecology, Lanzhou University, Lanzhou 730000
Gansu, P.R. China
2. International Yak Information Centre (IYIC), Department of Animal Science, Gansu Agricultural University, Lanzhou 730070, Gansu, P.R. China
To investigate genetic variation and differentiation in Chinese yak (Bos grunniens) populations, polymorphism at the β-Lactoglobulin (β-Lg) locus was examined in 608 yak milk samples, from six geographically isolated areas in Gansu Province, 39 milk samples of yak × cattle F1 hybrids and 24 milk samples of B. taurus cattle. Polyacrylamide gel electrophoresis detected six genotypes (AA, AB, AE, BB, BE and EE) and three alleles. The commonest allele in yak was allele E which frequency ranged from 1.00 (Tianzhu White yak) to 0.8049 (Xiahe yak). It is absent from cattle. The highest frequency at allele B, 0.1829, was observed in Xiahe yak. It is the commonest allele in the two cattle populations examined. Allele A was present in only two yak populations at very low frequency, 0.0068 (Luqu yak) and 0.0122 (Xiahe yak). Average genetic diversity within individual yak population (Hs) and within the combined total yak population (HT) was 4.8% and 5.2%, respectively. Average genetic differentiation (Gst) between yak populations was 8.4%. We assume that the presence of alleles A and B at the β-Lactoglobulin (β-Lg) locus in yak populations are the results of cattle introgression.
Keywords: Cattle, polyacrylamide gel electrophoresis, hybrid, milk, protein polymorphism, yak
The yak (Bos grunniens) is a unique animal, native to central Asian highland Himalayas, well adapted to the cold and high altitude environment. It is well known that domestic yak represent an important genetic resource for a large human population dwelling on the Himalayas, the Qinghai-Tibetan Plateau and other yak-rearing countries and areas neighbouring the Himalayas. At present, the estimated numbers of yak are over 14 million, of which more than 90% are found in China (Cai and Wiener 1995).
In dairy cattle, milk protein genes have been widely used as markers for increasing milk yield, altering milk composition and improving the physico-chemical properties of milk for the manufacture of various dairy products (Ng-Kawi-Hang 1998). Grosclaude et al. (1976a, b) were done studies on yak as early as 1976, studying milk protein variants in Nepalese yak. So far, four yak specific milk protein variants (β-LgE άs1 -CnE, άs2-CnC and k-CnX) have been characterised (Grosclaude et al. 1976a, b, 1982; Mahé and Grosclaude 1982; Kawamoto et al. 1992). In this study, we used yak β-Lactoglobulin (β-Lg) variants as a genetic marker to investigate the genetic variation within Chinese yak populations, differentiation between yak population and the level of cattle introgression in Chinese yak populations.
A total of 608 yak milk samples from six geographically isolated areas in Gansu Province were analysed. In addition 24 cattle (B. taurus) milk samples and 39 yak-cattle F1 hybrid milk samples were used as controls. The origin and number of milk samples examined in this study are listed in Table 1. Figure 1 shows the main distribution of yak in Gansu Province and it indicates the locations of the sampled populations.
Table 1. Populations sampled, origins and numbers (no.).
Populations |
Origins |
No. | |
Yak |
Maqu yak (MQ) |
Oula township, Maqu County |
190 |
Luqu yak (LQ) |
Along the provincial roads, Luqu County |
235 | |
Xiahe yak (XH) |
Sangke township, Xiahe County |
45 | |
Tianzhu White yak (TZW) |
Xidatan township, Tianzhu County |
21 | |
Tianzhu Black yak (TZB) |
Zhuaxixiulong township, Tianzhu County |
108 | |
Sunan yak (SN) |
Huangcheng Sheep Farm, Sunan County |
9 | |
Cattle |
Qingchuan cattle (QC) |
The vicinity of Linxia County |
10 |
China Holstein (CH) |
Dairy Farm of Gansu Agricultural University |
14 | |
Hybrid |
Yak × CH (F1) |
Datong Yak Farm, Qinghai Province |
35 |
Yak × local cattle (F1) |
Oula township, Maqu County |
4 | |
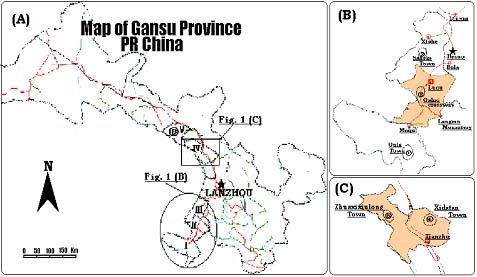
Note: (A) The main yak-rearing counties of Maqu, Luqu, Xiahe, Tianzhu and Sunan are represented as I, II, III, IV and V, respectively, where 1, 2, 3 in (B), 4, 5 in (C) and 6 in (A) show the detailed sampling sites for Maqu (MQ), Luqu (LQ), Xiahe (XH), Tianzhu White (TZW), Tianzhu Black (TZB) and Sunan (SN) yak, respectively.
Figure 1. Distribution of yak in Gansu Province.
Ten ml of middle-fresh milk from each animal was collected during the morning milking, frozen into liquid nitrogen and brought back to the laboratory for processing. Skimmed milk was prepared by centrifuging the milk samples at 1500 rpm for 15 minutes at room temperature, the upper milk fat was removed and the lower skimmed milk phase was kept at –20°C. The whey was separated using the acid precipitation method (Tsuji and Togamura 1987). The β-Lg variants were examined by discontinuous electrophoresis using a thin layer polyacrylamide gel system according to Erhardt (1993).
The values of HS, average gene diversity within populations, and HT, gene diversity in the combined yak populations, were calculated using GENEPOP (version 3.1). DST, average gene diversity between populations, and GST, relative magnitude of gene differentiation between populations, are defined as: DST = HT –HS and GsT = DST/HT, respectively (Nei 1987).
Genotypes and alleles frequencies are summarised in Table 2. We detected six genotypes at the β-Lactoglobulin locus named as AA, AB, AE, BB, BE and EE (Figure 2, Table 2).
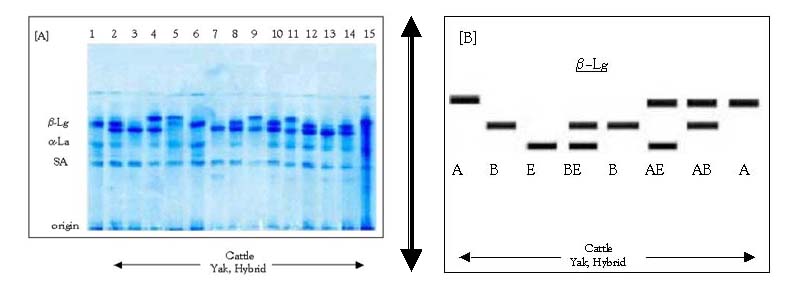
Note: (A) Simultaneous detection of á-Lactalbumin (á-La) and â-Lg by thin layer polyacrylamide gel electrophoresis (PAGE) (Erhardt 1993). All samples are of the genotype BB for á-La locus. Lanes 1, 6: b-Lg BB, Lanes 2, 8, 10, 12, 14: â-Lg BE, Lanes 3, 7, 13: â-Lg EE, Lanes 4, 9, 11: â-Lg AE, Lane 5: â-Lg AB, Lane 15: â-Lg AA. (B) Diagram showing the â-Lg variants as detected in (A).
Figure 2. The electrophoretic pattern of milk proteins around the b-Lactoglobulin (b-Lg) locus.
Table 2. Genotypes and alleles frequencies of β-Lg variations in yak, cattle and their F1 hybrids.
Locus |
MQ |
LQ |
XH |
TZW |
TZB |
SN |
F1 |
CH |
QC |
No. = 181 |
No. = 221 |
No. = 41 |
No. = 16 |
No. = 89 |
No. = 8 |
No. = 39 |
No. = 13 |
No. = 10 | |
|
Genotypes β-Lg |
|||||||||
AA |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
1 |
1 |
AB |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
8 |
6 |
AE |
0 |
3 |
1 |
0 |
0 |
0 |
9 |
0 |
0 |
BB |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
4 |
3 |
BE |
2 |
61 |
14 |
0 |
3 |
2 |
30 |
0 |
0 |
EE |
179 |
157 |
26 |
16 |
86 |
6 |
0 |
0 |
0 |
| Alleles | |||||||||
| ß-LgA | 0 | 0.0068 | 0.0122 | 0 | 0 | 0 | 0.1154 | 0.3846 | 0.4000 |
| ß-LgB | 0.0055 | 0.1357 | 0.1829 | 0 | 0.0169 | 0.1250 | 0.3846 | 0.6154 | 0.6000 |
| ß- LgE | 0.9945 | 0.8575 | 0.8049 | 1.0 | 0.9831 | 0.8750 | 0.5000 | 0 | 0 |
| Note: MQ = Maqu; LQ = Luqu; XH = Xiahe; TZW = Tianzhu White; TZB = Tianzhu Black; SN = Sunan yak. | |||||||||
The Tianzhu White yak population was monomorphic showing only the EE pattern. Polymorphisms were found in the other yak populations, in cattle and the F1 hybrids with the eventual presence at the A, B and E alleles. Allele E was not detected in cattle. It is worth mentioning that alleles A and B, which were previously only observed in B. taurus cattle (Kawamoto et al. 1996), are detected exclusively in the heterozygous forms (AE and BE) in yak and their F1 hybrids. Similarly, allele E, which was proved to occur only in yak (Grosclaude 1976a; Bell et al. 1981a, b) and Bali cattle (Bell et al. 1981a, b), is also only occurring in heterozygote forms (AE and BE) in F1 hybrids.
The frequency of allele E (not including Tianzhu White yak where it is fixed) is the highest in Maqu yak (0.9945) followed by Sunan yak (0.9831), and the lowest in Xiahe yak (0.8049). The frequency of allele B is the highest in Xiahe yak (0.1829) followed by Luqu yak (0.1357) and the lowest is in Maqu yak (0.0055). Allele A is only present in two populations of yak, Luqu (0.0068) and Xiahe yak (0.0122). The frequency of allele E in F1 hybrids is 0.5; it suggests that the parental yak population was fixed for this allele (Table 2). Similar results were obtained in F1 hybrids by (Grosclaude et al. 1976a, Zhang et al. 1991).
Genetic variation within yak populations were estimated by Nei’s genetic diversity coefficients (Nei 1987) using three milk protein loci of a-Lactalbumin (a-La), b-Lactoglobulin (b-Lg) and b-Casein (b-Cn) (Table 3).
Table 3. Genetic diversity (Hs, Ht) of three milk protein loci.
| MQ | LQ | XH | TZW | TZB | SN | |||
| Locus | No. = 180 | No. = 220 | No. = 41–44 | No. = 16 | No. = 89 | No. = 8–9 | Mean3 | Pooled4 |
| a-La1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| ß-Lg | 0.0110 | 0.2463 | 0.3186 | 0 | 0.0331 | 0.2188 | 0.1380 | 0.1432 |
| ß-Cn2 | 0 | 0.0357 | 0 | 0 | 0 | 0 | 0.0060 | 0.0139 |
| Mean | 0.0037 | 0.0940 | 0.1062 | 0 | 0.0110 | 0.0729 | 0.0480 | 0.0524 |
| 1, 2: data according to Qi (2000);
3: HS , 4: HT. MQ = Maqu; LQ = Luqu; XH = Xiahe; TZW = Tianzhu White; TZB = Tianzhu Black; SN = Sunan yak |
||||||||
No genetic variations are found in Tianzhu White yak, whereas, in another two neighbouring populations (Figure 1, A and C), Tianzhu Black yak (He = 1.1%) and Sunan yak (He = 7.29%) the expected heterozygosity is low. For the Maqu, Luqu and Xiahe yak populations, which originated on Gannan highland pastures and with bordering geographic areas (Figure 1, B), genetic variations were detected in all populations ranging from 0.37% (Maqu yak) to 10.62% (Xiahe yak). Although genetic variations exist within yak populations, the average gene diversity within (HS = 4.8%) and between populations (DST = 0.44%) is very small, and so is the genetic diversity grouping all yak population studied in a single one (HT = 5.2%). The average gene differentiation among yak populations (GST) of 8.4% indicates that population differentiation is low at these three milk protein loci.
It is generally accepted that domestic yak were descended from wild yak (B. mutus) caught and tamed by ancient Qiang people (a collective name for all ancient nomads in western China) in the Qiangtang and other areas of northern Tibet, and subsequently distributed to the current areas where yak are found (Zhang 1989; Cai and Wiener 1995). Out of the 18 China yak breeds, 10 are widely recognised (Zheng 1985). Although these different yak breeds, for the most part, are living in different areas, they are sometimes sharing grazing habitats. The natural grazing and transhumance practices make genetic exchange between populations feasible. So, considering its likely single origin and the grazing systems followed by the nomads, all yak populations or breeds might have relatively uniform genetic backgrounds. It is the case for the Nepalese yak (Grosclaude et al. 1976a, Grosclaude et al. 1976b; Kawamoto et al. 1992), the Mongolian yak (Grosclaude et al. 1982) and most of Chinese yak populations (Qi 2000) at the level of milk protein loci. Also, it has been observed that yak populations are sharing the same allele at blood protein and enzyme loci (Men et al. 1989; Namikawa et al. 1992; Tu 1996).
B. taurus cattle breeds have been introduced to yak-raising areas in order to produce hybrids (Zhang 1989; Cai 1990). In areas with better communication systems (e.g. Luqu County) as well as in agropastoral areas (e.g. Xiahe County), such breeding practices are easier to implement. As a result, yak in these areas are more often crossbreed with local cattle and exotic western cattle breeds resulting in a change of their genetic background. Although such changes are probably minor, it would explain the presence of some low frequency alleles of cattle origins likely in these populations.
The authors thank the International Yak Information Center (IYIC) for financial support. Mr Tao Shixin, Prof Dr Du Guozhen, Mr Yang Xichao, Mr Liang Yulin and Mr Hu Jiang provided invaluable assistance to milk sampling in the field.
Bell K., McKenzie H.A. and Shaw D.C. 1981b. Bovine ß-lactoglobulin E, F and G of Bali, Banteng, Cattle, Bos, Bibos, javanicus. Australian Journal of Biological Science 34:133–147.
Cai L. 1990. Sichuan yak. Sichuan Ethnic Press, Chengdu, P.R. China. 223 pp. [in Chinese].
Cai L. and Wiener G. 1995. The yak. FAO (Food and Agriculture Organization of the United Nations) Regional Office for Asia and the Pacific, Bangkok, Thailand. 237 pp.
Erhardt G. 1993. A new αs1-casein allele in bovine milk and its occurrence in different breeds. Animal Genetics 24:65–66.
Grosclaude F., Mahé M.F., Mercier J.C., Bonnemaire J. and Teissier J.H. 1976a. Polymorphisme des lactoprotéines de bovinés népalais. I. Mise en évidence, chez le yak, et caractérisation biochimique de deux nouveaux variants: ß-lactoglobuline Dyak et caséine αs1E. Annales de Génétique et de Sélection Animale 8(4):461–479.
Grosclaude F., Mahé M.F., Mercier J.C., Bonnemaire J. and Teissier J.H. 1976b. Polymorphisme des lactoprotéines de bovinés népalais. II. Polymorphisme des caséines as-mineures; le locus αs2-Cn est-il lié aux loci αs1-Cn, ß-Cn et -κCn? Annales de Génétique et de Sélection Animale 8(4):481–491.
Grosclaude F., Mahé M.F. and Accolas J.P. 1982. Note sur le polymorphisme génétique des lactoprotéines de bovins et de yak Mongols. Annales de Génétique et de Sélection Animale 14(4):545–550.
Kawamoto Y., Namikawa T., Adachi A., Amano T., Shotake T., Nishida T., Hayashi Y., Kattel B. and Rajubhandary H.B. 1992. A population genetic study on yak, cattle and their hybrids in Nepal using milk protein variations. Animal Science and Technology (Japan) 63(6):563–575.
Kawamoto Y., Hongo A., Inamura T. and Shrestha H.R. 1996. An additional survey of gene flow between yak and cattle in the Nepalese Himalayas. Animal Science and Technology (Japan) 67(4):373–377.
Mahé M.F. and Grosclaude F. 1982. Polymorphisme de la caséine as2 des Bovinés: caractérisation du variant C du Yak (Bos grunniens). Annales de Génétique et de Sélection Animale 14(4):401–416.
Men Zhenming, Han Jianlin and Zhang Yuezhou. 1989. Polymorphisms of blood proteins in Tianzhu white yak. Journal of China Yak 4:3–5.
Namikawa T., Amano T., Yamamoto Y., Tsunoda K., Shotake T., Nishida T. and Rajbhandary H.B. 1992. Genetic differentiation in blood groups and blood protein variations of Nepalese cattle, yak and zoopa. Report of the Society for Resarch on Native Livestock (Japan) 14:17–37.
Nei M. 1987. Genetic variation within species. In: Nei M. (ed), Molecular evolutionary genetics. Columbia University Press, New York, USA. pp. 177–207.
Ng-Kawi-Hang K.F. 1998. Genetic polymorphism of milk proteins: Relationships with production traits, milk composition and technological properties. Canadian Journal of Animal Science 78:131–147.
Qi Xuebin. 2000. Genetic variations of yak in Gansu: Inferred from their milk protein polymorphisms. MSc thesis. Gansu Agricultural University, Lanzhou, P.R. China. 31 pp. [in Chinese].
Tsuji S. and Togamura H. 1987. An introduction of new method for simultaneous detection of milk protein variants (in Japanese). Animal Blood Groups Research Information (ABRI) 15: 17–21.
Tu Zhenchang. 1996. Genetic diversity and genetic divergence in Chinese yak populations. PhD thesis. Northwestern Agricultural University, Yangling, P.R. China. 62 pp. [in Chinese].
Zhang C.J., Fang Z.L., Li Q.J. and Zhang S. 1991. Polymorphisms of ß-Lactoglobulin in cattle–yak hybrids and their parents. Journal of China Yak 1:18–22.
Zhang Rongchang. 1989. China: The yak. Gansu Scientific and Technology Press, Lanzhou, P.R. China. 386 pp. [in Chinese].
Zheng P.L. (ed). 1985. Chinese animal breeds and their ecological characteristics. Agricultural Press, Beijing, P.R. China. pp. 45–59. [in Chinese].